
ITHILDIN
Integrated Tool for Homologous Image Landmark Detection in Insects using Neural networks
This tool is designed for Diptera identification and geometric morphometric analysis based on uploaded images of wings. We refer to this preprint for further information. The system uses AI to annotate landmark and semilandmarks on the vein patterns of the wings of Diptera. Currently it is trained for Culicidae, Drosophila and Glossina. In addition we utilise the CNN provided by BALROG to predict the species of mosquito. BALROG is trained on 34 mosquito species while some species are summarized under the label "other". You will receive a unique identifier for each submission for future reference, please use it if you contact us regarding issues.
Search Previous Results
Already processed images? Enter your session ID to retrieve your results.
Upload Files
For optimal prediction, please ensure that the images have a neutral background and clearly show a single, intact wing. We highly recommend reviewing our Image Capture Guidelines for detailed instructions on preparing your wing specimens. Acceptable file formats include JPG, JPEG, PNG, TIF and TIFF and do not use "." as part of the file name. If you upload other files they will be ignored. You can upload up to 256 MB per upload. For regular uploads or large amounts of images we recommend to install the application locally. You can find the repository and installation instructions here. The model takes on average 3 to 5 seconds per image to classify. Please confirm that you own or have the necessary rights for the images you upload. Click here to get an example prediction.
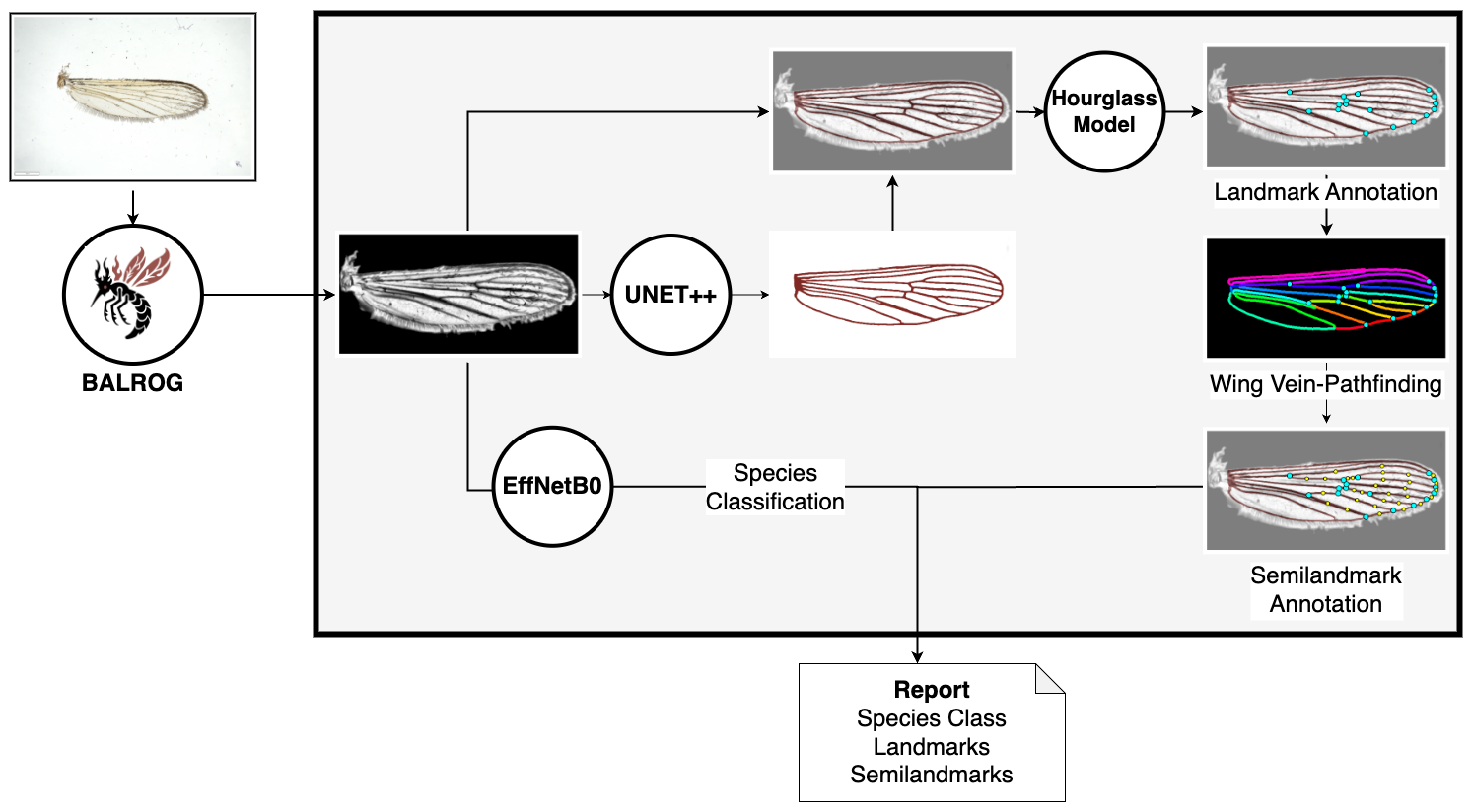
Note: Species identification is only available for mosquito wings. The models for drosophila and tsetse are intended for demonstration only. Due to training on a small number of samples, their predictions may be unreliable and should be interpreted with caution.
Contact Us
K. Nolte, F. G. Sauer, R. LühkenBernhard-Nocht Institute for Tropical Medicine
Bernhard-Nocht-Straße 74
20359 Hamburg, Germany
